The paper titled "Channeling Fisher: Randomization Tests and the Statistical Insignificance of Seemingly Significant Experimental Results" (.htm) is currently the most cited 2019 article in the Quarterly Journal of Economics (372 Google cites). It delivers bad news to economists running experiments: their p-values are wrong. To get correct p-values, the article explains, they need to run, instead of regressions, something called "randomization tests" (I briefly describe randomization tests in the next subsection). For example, the QJE article reads:
"[the results] show the clear advantages of randomization inference . . . [it] is superior to all other methods" (p.585).
In this post I show that this conclusion only holds when relying on an unfortunate default setting in Stata. In contrast, when regression results are computed using the default setting in R [1], a setting that's also available in Stata, a setting shown over 20 years ago to be more appropriate than the one used by Stata (Long & Ervin 2000 .htm), the supposed superiority of the randomization test goes away.
Randomization is not superior, Stata's default is inferior.
In what follows I briefly explain randomization tests, then briefly explain the need for robust standard errors. Then I describe my re-analysis of the QJE paper.
All scripts and files for this post are available from https://researchbox.org/431
Randomization Tests
Randomization tests are often attributed to Ronald Fisher, which is why the QJE paper's title begins with "Channeling Fisher", but this footnote explains why that may not be quite right [2].
Let's get a sense of how randomization tests work.
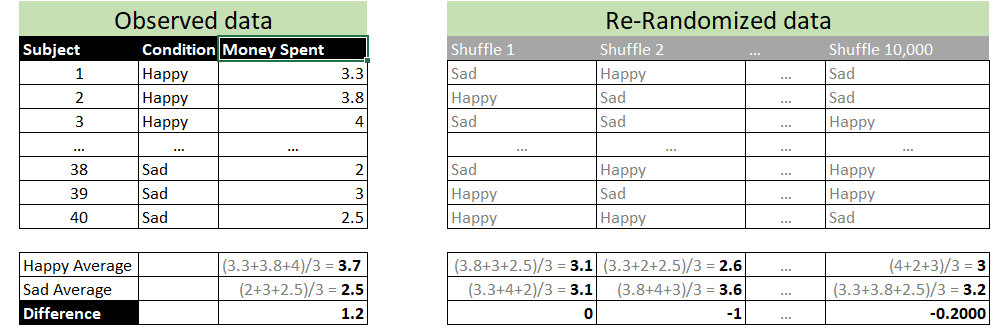
Say that in an experiment, participants are put in a happy vs sad mood and then they go shopping. Happy participants spend $1.20 more on average. To compute a p-value for this mean difference we could use a t-test, but we could also use a randomization test. For the latter, you randomly shuffle the condition column (Happy vs. Sad); see the gray columns in the table on the right above. A datapoint from a participant in the happy condition can be re-labeled as sad, or stay happy, at random. You then compute the mean difference between the shuffled "happy" vs "sad" conditions. For example, the first shuffle leads "happy" and "sad" participants to, coincidentally, have means of $3.10, for a mean difference of $0. You repeat this shuffling process, say, 10,000 times. You then ask: what % of the 10,000 shuffled datasets obtained a mean difference of $1.20 or more? That's your randomization test p-value [3].
Robust Standard Errors
Many readers know that "heteroskedasticity" describes something that can go wrong with a regression. In a nutshell, the problem is that the regression formulas assume that all observations are equally noisy, but in some situations some observations can be systematically noisier than others, creating a problem. To intuitively think of heteroskedasticity let's think about wine and academics. When professors buy a bottle of wine, sometimes they spend a lot, and sometimes they spend little. When PhD students buy a bottle of wine, they always spend little. Professors' wine spending is thus systematically noisier than student wine spending. So wine spending is heteroskedastic.
With heteroskedasticity, regression standard errors tend to be wrong, and thus p-values tend to be wrong.
The solution to incorrectly assuming all observations are equally noisy is to not assume that all observations are equally noisy, and to instead estimate how noisy each observation is. This is done by calculating robust standard errors, which are robust to heteroskedasticity.
Robust standard errors were developed surprisingly late in the history of statistics. White (1980) had the first major insight, and then MacKinnon & White (1985) developed what are today's most popular approaches to computing robust standard errors (See MacKinnon's (2013) review of the first 30 years of robust & clustered SE).
Note: the previous paragraph was added two weeks after this post was first published (also switching STATA->Stata). See the original version (.htm).
In any case, there are now five main ways to compute robust standard errors.
Stata has a default way, and it is not the best way.
In other words, when in Stata you run: reg y x, robust you are not actually getting the most robust results available. Instead, you are doing something we have known to be not quite good enough since the first George W Bush administration (the in-hindsight good old days).
The five approaches for computing robust standard errors are unhelpfully referred to as HC0, HC1, HC2, HC3, and HC4. Stata's default is HC1. That is the procedure the QJE article used to conclude regression results are inferior to randomization tests, but HC1 is known to perform poorly with 'small' samples. Long & Ervin 2000 unambiguously write "When N<250 . . . HC3 should be used". A third of the simulations in the QJE paper have N=20, another third N=200.
So, the relevant question to ask during the (first) Biden administration is whether randomization is better than HC3 (Narrator: it is not).
Re-doing the simulations in the QJE paper
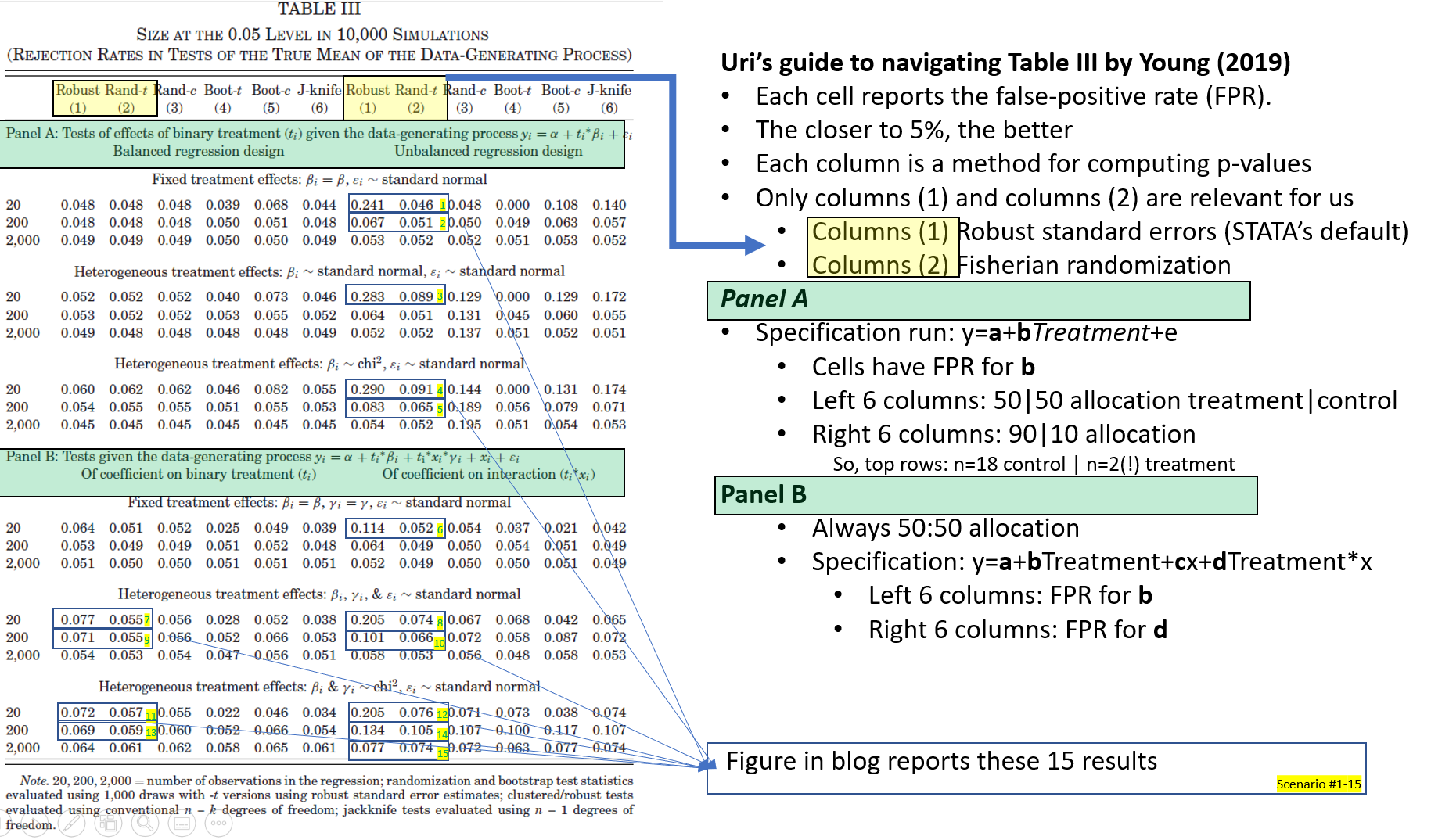
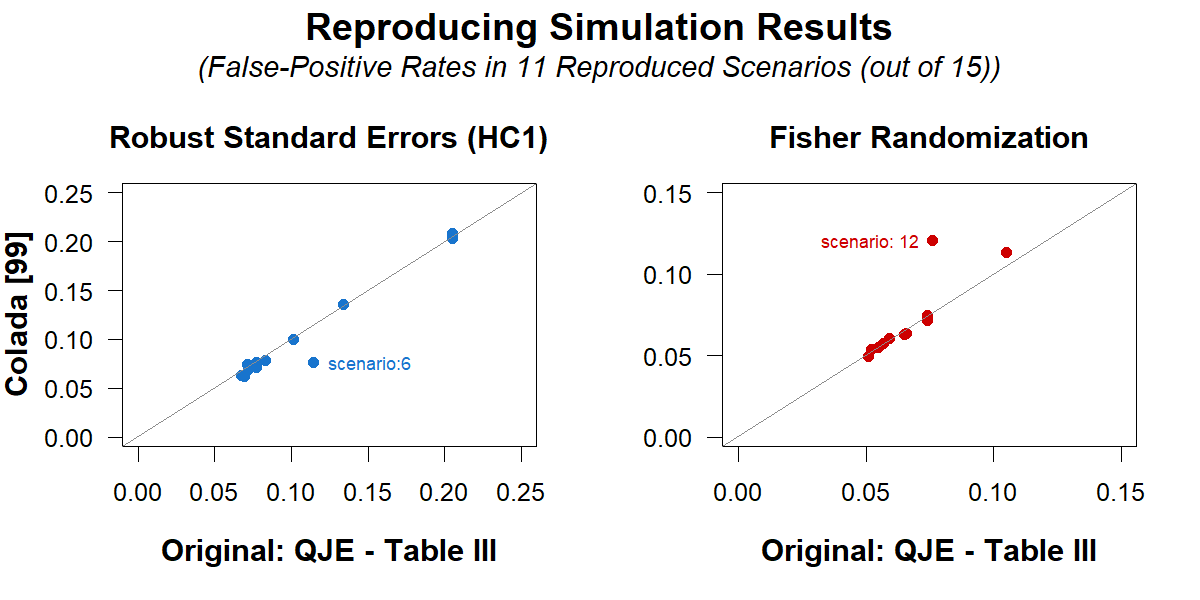
The QJE paper compares the performance of robust standard errors (HC1, i.e., Stata's default) with randomization tests for a variety of scenarios (involving various degrees of heteroskedasticity), by computing false-positive rates, that is, computing how often p<.05 is observed when the null is true [4]. The paper reports that regression results have false-positive rates that are too high, rejecting the null up to 21% of the time instead of the nominal 5%, while the randomization tests perform much better.
I think that the simulated scenarios in the QJE paper are too extremely heteroskedastic, making Stata's default look worse than it probably performs on real data, but I wanted to rely on the same scenarios to make it easy to compare my results with those in the QJE paper. Plus, I don't really know how much heteroskedasticity to expect in the real world.
I began by successfully reproducing in R the reported simulation results (Table III of the QJE paper). Because I am focusing on the subset of scenarios for which regressions had an inflated false-positive rate (>6.5%), I made an annotated version of that Table III, that makes it clear which results I am reproducing. Most readers do not need to see it, but you can if you want
After reproducing the published results, I re-ran the analyses using robuster standard errors (HC3). The differences between blue and red bars reproduce the QJE results: Stata's default is worse than randomization. This wraps up the point about randomization tests. The QJE paper also re-analyzes data from 53 economics experiments and concludes "one of the central characteristics of my sample [of economics experiments] is its remarkable sensitivity to outliers" (p.566), adding "With the removal of just one observation, 35% of .01-significant reported results in the average paper can be rendered insignificant at that level." (p.567). I didn't have intuitions as to whether these numbers were actually troubling, so I looked into it. I don't think they are actually troubling. That proportion, 35%, is in the ballpark of what to expect when there are zero outliers. I wrote about this, but then after discovering the HC1 vs HC3 thing, worried it is too niche and makes the post too long, so I hid it behind the button below.
Take home To produce digestible and correct empirical analyses we need to decide what is and what is not worth worrying about. For most papers, randomization tests are not worth the trouble. But remember, when doing robust standard errors in Stata, either with experimental or non-experimental data: Author feedback
References Footnotes.
All results are shown in the figure below [5].
The (mostly) lack of difference between the red and gray bars show that this is because Stata's default is especially bad, rather than because randomization tests are especially good. If anything, robuster standard errors outperform randomization tests in these scenarios.
I am a fan of randomization tests. They are useful when no accepted statistical test exists (e.g., to compare maxima instead of means), they are useful to run tests on complex data structures (e.g., we use it our Specification Curve paper .pdf to test jointly many alternative specifications run on the same data), they are useful for correcting for multiple comparisons with correlated data, they are useful for testing unusual null hypotheses (e.g., in fraud detection). But, if you are analyzing data with random assignment, in anything larger than a laughably small sample (n>5), why bother with a procedure that needs to be coded by hand (and is thus error prone), that is hard to adjust to complex designs (e.g., nested designs or stratified samples), and that does not deliver confidence intervals or other measures of precision around the estimates?![]()
Our policy (.htm) is to share drafts of blog posts that discuss someone else's work with them, to solicit suggestions for things we should change prior to posting, and to invite a response that is linked to from the end of the post (i.e., from here). I shared a draft of this post with Alwyn Young on September 30th, and then sent a reminder on October 9th, but he has not yet replied to either message. We did exchange a few emails prior to that, starting with an email I sent on September 9th, 2021, but that discussion did not involve the HC1 vs HC3 issue. I should probably point out that we have had n=99 blog posts so far, and removing a single observation, 100% of authors have replied to our emails requesting feedback.
sandwich .htm [↩]
The scatterplot from this footnote, and associated calculations, are done in Section #7 in the R script available here: https://researchbox.org/431.13 [↩]