About a year ago I wrote Colada[95], a post on the threat R poses to reproducible research. The core issue is the 'packages'. When using R, you can run library(some_package) and R can all of a sudden scrape a website, cluster standard errors, maybe even help you levitate. The problem is that packages get updated often, and on occasion in 'backwards incompatible' ways, making your existing code obsolete. The code that works today, may not work tomorrow.
In that post I introduced a solution to this problem with R packages: a new R package. It's called groundhog.
With groundhog, the only thing you need to change to make your R code reproducible is:
Instead of: library(pkg)
Do this: groundhog.library(pkg, date)
Now every time you run that code, you load the version of the package that was available on that date, regardless of when you run it. That's really all you need to do to dramatically improve the reproducibility of your R Code.
This post is an update on three fronts:
1) Share new evidence on iRreproducibility (geRit?)
2) Announce groundhog 2.0 (key new feature: it works with GitHub packages)
3) A 'show me the money' moment.
If you are familiar with renv and wonder how it compares to groundhog, check this out (.htm)
1) New evidence that R is a threat to research reproducibility
Over the past year I have learned a few things that made me more concerned about R's reproducibility (more concerned than when I was motivated to spend months of my life developing a fricking R package to address that concern). I will mention three things.
Thing #1: Most posted R scripts apparently don't even run
A paper published a few weeks ago in Nature: Scientific Data (.htm) attempted to automatically re-execute 2335 R scripts posted as supporting materials for published papers. After cleaning the scripts (installing necessary packages and fixing paths to local files) only 44% of scripts run without generating errors. So, most scripts did not run. Moreover, 21% of all failures were attributed to packages not loading. This is an underestimate of the problem caused by packages, because a package may load successfully, but reproduce different results, or produce an error elsewhere (see "show me the money" example at the end of this post).
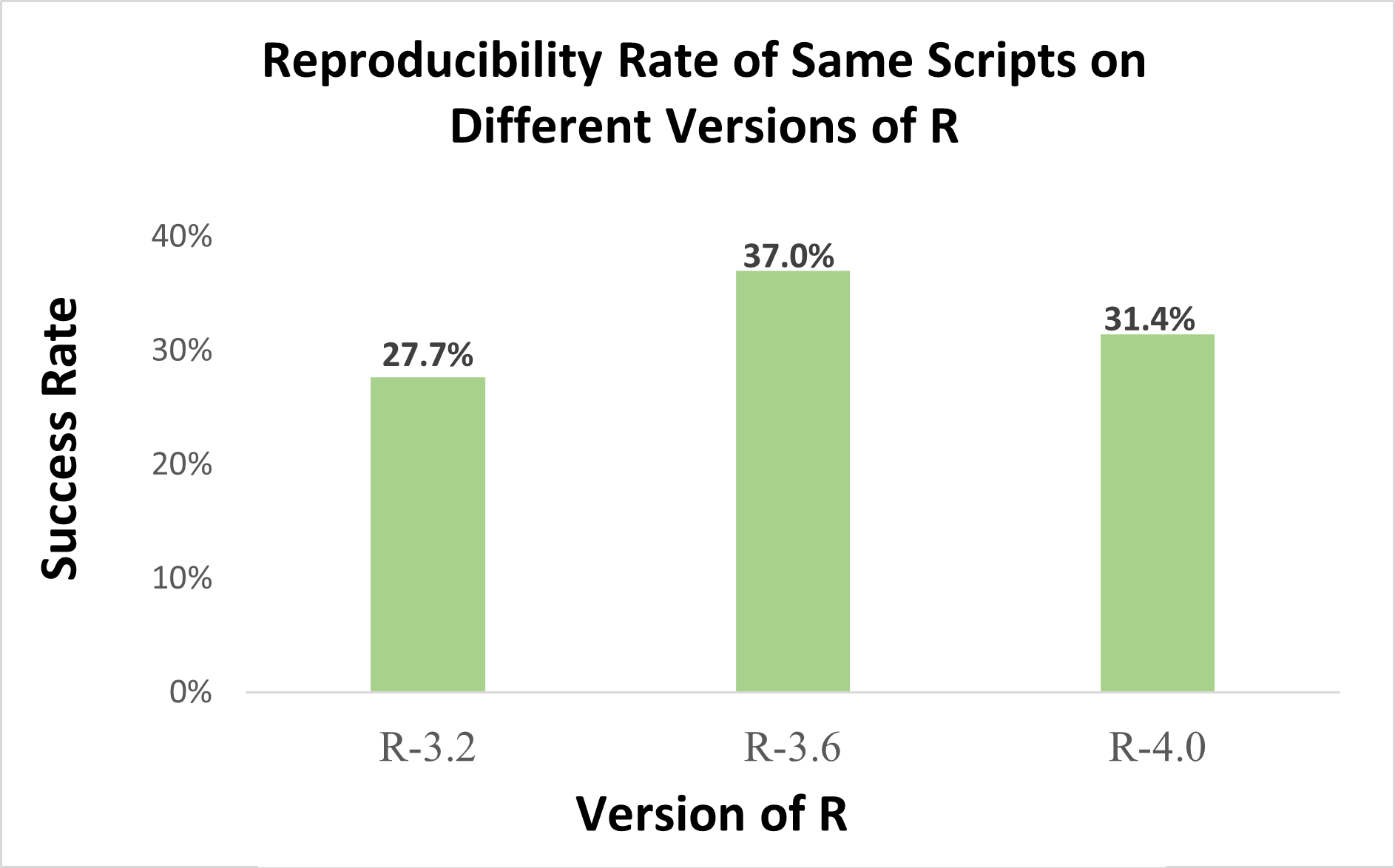
The script cleaning was done in an automated way, and this is important. It seems likely that some of the scripts that did not run, could run if a person were reading and editing the code before executing it. But there is one key finding that is free of this ambiguity: the same scripts run in some but not other versions of R.
Fig 1. Uri made this figure based on numbers reported in Figure 11 (.png) of Trisovic et al. (2022).
R-3.2 may feel like ancient history, but when Trump was running for president, R was running 3.2.
Groundhog also helps with this source of irreproducibility. Specifically, when the date entered in groundhog.library() does not match the R version being used, it gives a warning and suggests a date to switch the groundhog day to, or the version of R to use for that groundhog day.
Running multiple versions of R on the same computer is trivial in Windows and quite easy in a Mac. Groundhog's website provides step-by-step instructions.
Thing 2. Come think of it: this is a big pain in the Rs for books
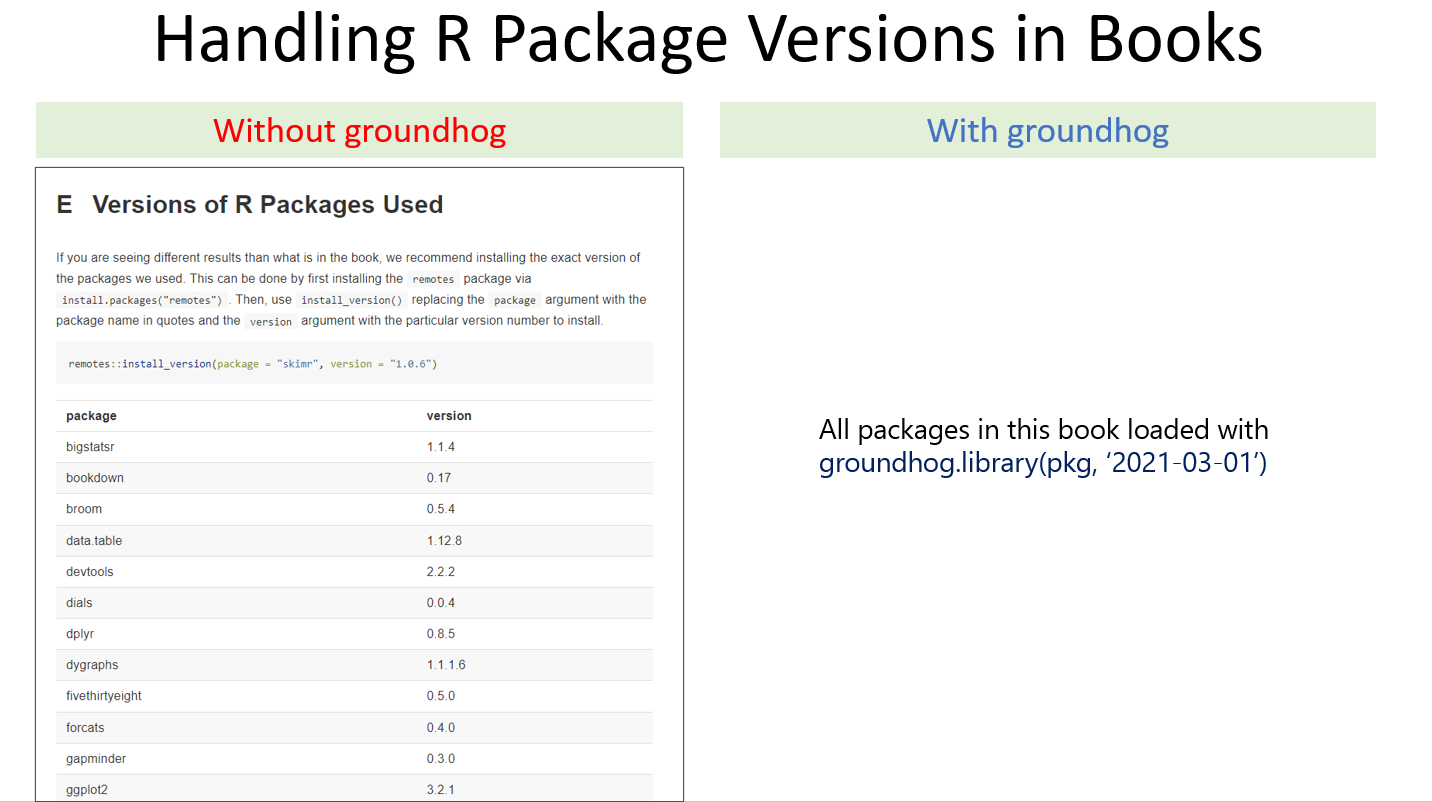
I recently noticed that books about R, or that rely on R, often include printed out lists of the version of each package they used (see figure below). This highlights that I am not the only one worried about the issue of R stability. And just think about it. You may work for years on a book on causal inference, say, or interpreting interactions, and the examples you include in your book, if they involve R code, may stop working shortly after the book is finished (possibly even before it is published). This seems tolerable for a book about R, say "R for Data Science" (as planned obsolescence is not a terrible business model), but it seems less tolerable for a book on anything else that happens to use R to give concrete examples. A book on causal inference is not going to be revised every 6 months just to make sure all the R examples still run. In fact, it may never get revised.
Attempting to fix a book's reproducibility problem by printing a list of used package versions is both impractical and dangerous (see footnote for why it is impractical and dangerous [1]). An alternative is to write books relying on groundhog.
In fact, textbooks provide a textbook example of groundhog's simplicity:
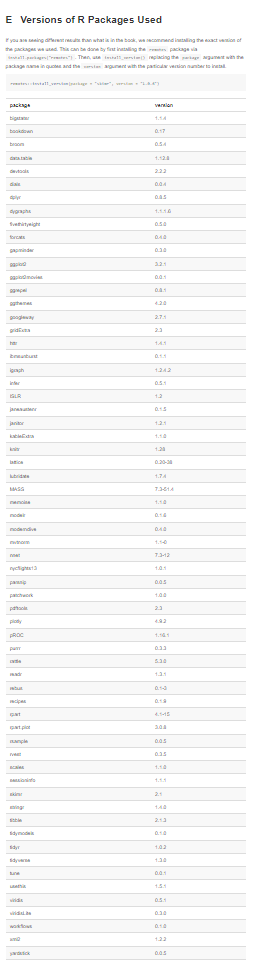
That figure in the left is actually a small portion of the full list. Check. It. Out.:
Thing 3. Personal Experience with abandoned packages
While working on a research project I needed to revisit my own code from just a few months earlier. But, I could not install a package that I needed because it was no longer available on CRAN (it had been archived because one of its dependencies was no longer being maintained). My own code would not have run in my own machine. But I had groundhog, and so it did.
2. New features since groundhog 1.0.0
The current version of groundhog on CRAN is v1.5.0. It has some neat features introduced since the original release v1.0.0, including the possibility of loading/installing a set of packages in a single call:
pkgs <- c('metafor','pwr', 'jsonlite')
groundhog.library(pkgs,'2022-03-01')
And having version-control for groundhog itself
meta.groundhog('2022-01-01') #load the version of groundhog available that date.
A bigger innovation is that the next release of groundhog, v2.0.0, will work not just with CRAN but also with git repositories (GitHub and GitLab).
Like this:
groundhog.library('crsh/papaja' , '2022-03-01')
groundhog.library('gitlab::jimhester/covr', '2022-03-01')
Git packages arguably need version control even more than CRAN packages do; see footnote [2]
Groundhog 2.0.0 should be on CRAN by May 2022.
In the meantime, you can use the almost there version: v1.9.9.9999 available on GitHub:
remotes::install_github('CredibilityLab/groundhog')
3) Show me the money
I thought it would be interesting to try and see if groundhog could 'rescue' an R script that the Nature: Scientific Data paper reported as failing to reproduce.
I downloaded one R script flagged as non-reproducible, supporting a paper published in Political Analysis (.htm) [3]. The script has just 9 active lines of code; it is supposed to estimate a regression with fixed effects using the package 'bife' (after loading the data with 'foreign').
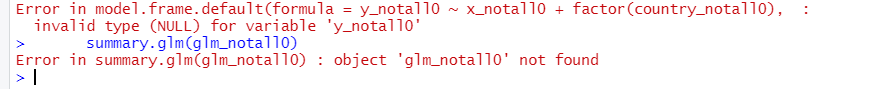
After running that script in R-4.1.3 (current version as of this writing), however, all I got was this ugly thing:
The R script had been uploaded to the Dataverse on January 2019, so I thought to try late 2018 as a reference date. Back then R was running version R-3.5. So I started up R-3.5 in R Studio (again, it is easy to have multiple versions of R in the same computer; see how).
I then did install.packages(c('foreign','bife')), loaded both packages with library() and ran the 9 lines again. And…
…no luck. Got the same error message.
So I used groundhog.libray() instead.And …
…that did work.
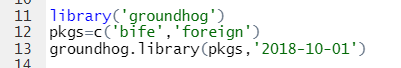
Specifically, I installed & loaded the packages like this:
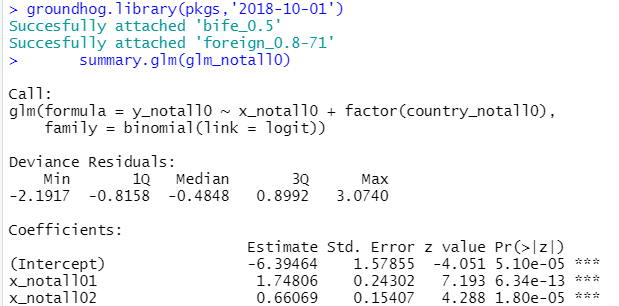
And the 9 lines of code produced this beautiful, oddly self-affirming, regression table:
If curious, this is the R Code for the above example.
See footnote for why the code ran with groundhog.library() but not with library() [4].
Conclusions
A big lesson from Bill Murray's conundrum in the movie Groundhog Day, is that if you want different results, you need to try something different. We know by now that relying on R alone for package management (install.packages() + library()) will lead researchers to share R scripts that do not work. So if we want the R scripts we share to work we need to try something different. Groundhog is something different. Nothing is perfect, but groundhog delivers a huge increase in reproducibility with absolutely minimal effort.
groundhog.library(pkg, date)
Learn more about groundhog:
Colada[95] on Groundhog | groundhogr.com | GitHub
![]()
Author feedback
I shared an early draft of this post with Ana Trisovic (.htm), 1st author of the Nature: Scientific Data paper and she provided useful feedback and clarifications. The post did change substantially since she read it.
Footnotes.
- Impractical because the kind of person who needs an R book will probably struggle understanding what that's all about or how to handle the list. Dangerous because if a package does change, and does break an example in the book, simply installing the old version will, OK, maybe fix this book, but probably break other scripts written based on the newer version of the package (e.g., examples in newer books). [↩]
- Git packages need version control even more than CRAN packages for two opposing reasons. First, some git packages get edited very frequently, some daily, but their version numbers do not get updated; thus the same package-version is actually a different package on Monday vs Tuesday. With groundhog packages are identified by date, so that's fixed. Second, and on the other hand, some packages are updated very infrequently. Which means they can become unusable when the CRAN packages they depend on get updated but the git package does not. Loading git packages with groundhog, you will always load the dependencies that package actually needs and relied on when you first wrote your script. Having said that, it is not possible to version control git packages as reliably as CRAN packages. See why here: (.htm [↩]
- I had two attempts prior to this one. Both actually ran as-is, so I did not reproduce their irreproducibility; in these cases there was no room for groundhog to improve anything. The third script I tried did produce an error. That's the one discussed above. [↩]
- When you run
install.packages()in R, you get whatever version of the package happens to be the most recently available on CRAN (for the version of R that you are running). For the package 'bife', CRAN's most recent version of bife, for R-3.5.3, isbife_0.7, which was released nearly a year after the Political Analysis paper was published; back then the current version wasbife_0.5. So, runninginstall.packages()in 2022 in R-3.5.3, I getbife_0.7, but I needed versionbife_0.5, the version loaded bygroundhog[↩]